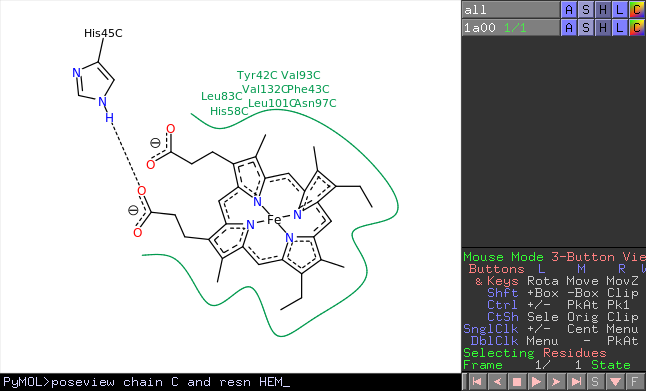
Understanding representations of the molecular world is critical to becoming an expert in the biomolecular sciences 1, because the interpretation of such images is key to understanding biological function 2.

The enzyme is bound to one of its substrates, as well as a non-reactive substrate analog, which allows the user to analyze interactions in the catalytic complex. The model chosen for this protocol is human glucokinase, an isoform of the enzyme hexokinase, which catalyzes the first step of glycolysis.
#PYMOL TUTORIAL HOW TO SHOW LIGAND INTERACTIONS FREE#
The protocol enables the user to model an active site using a specific visualization program, or to sample several of the free programs available. This guide is intended for students seeking to learn the basics of a specific program, as well as instructors incorporating biomolecular modeling into their curriculum. In this protocol, we describe this process using four freely available macromolecular modeling programs: iCn3D, Jmol/JSmol, PyMOL, and UCSF ChimeraX. A critical skill in this area is to model a protein active site, displaying parts of the macromolecule that can interact with a small molecule, or ligand, in a way that shows binding interactions.
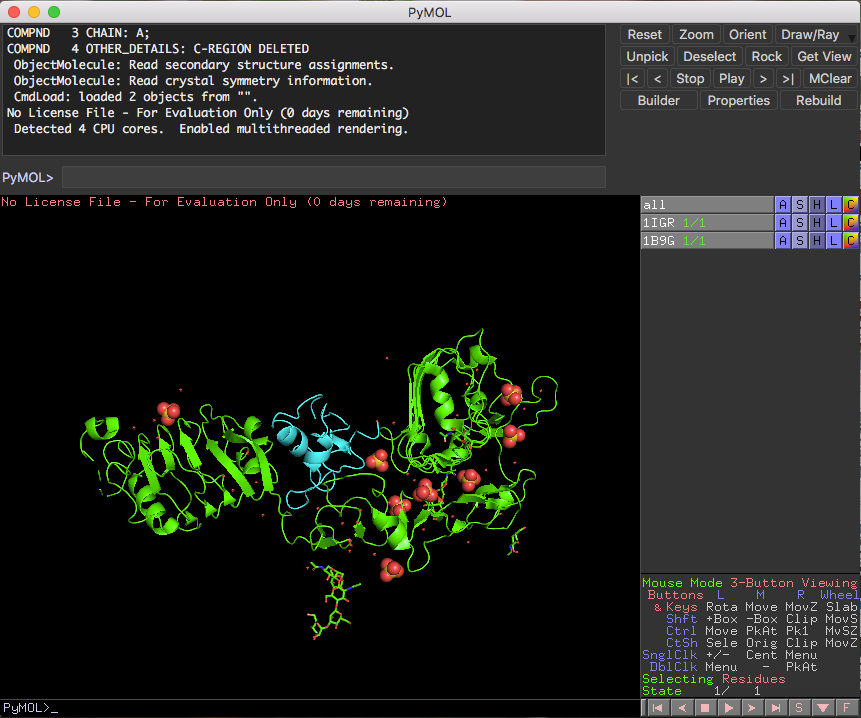
Various programs allow a learner to manipulate 3D structures, and biomolecular modeling promotes active learning, builds computational skills, and bridges the gap between two dimensional textbook images and the three dimensions of life. Biomolecular visualization skills are paramount to understanding key concepts in the biological sciences, such as structure-function relationships and molecular interactions.
